A New Paradigm in Bacterial Transcriptional Regulation
A previously unknown mechanism relies on intrinsically inactive σ factor
In collaboration with researchers from SYNMIKRO in Marburg, Max Planck scientists have described a previously unrecognized regulatory mechanism in bacterial signal transduction and transcriptional regulation, which seems to be widespread in bacteria.
Introduction
Within their environment, bacteria often encounter various kinds of changes. In order to survive under stress or changing conditions, bacteria have to respond fast and adequate to achieve a physiological adaptive response. This mostly happens by specifically adapting gene expression. Thus, transcriptional regulation is one of the biggest means by which bacteria adapt to external stress conditions.
A novel mechanism of transcriptional regulation
Initiation of transcription in bacteria only occurs upon the binding of a key component known as the sigma factor (σ factor) to the RNA polymerase (RNAP) core enzyme in order to form the holozyme. This holozyme then recognizes key promoter elements and subsequently, enables transcription. During external stress conditions, the primary σ factor is replaced by an alternative σ factor, which differs from the former with respect to the promoter sequences that it recognizes. Upon association of this alternative σ factor with the RNAP core enzyme in formation of the holozyme, transcription of the corresponding stress-response genes occurs.
Among the various classes of alternative sigma factors, the most abundant are the extracytoplasmic function (ECF) σ factors. Sigma factors are most commonly known to be intrinsically active, which means that the bacterial cell has to keep them in an inactive state until their action is warranted. There are several mechanisms through which sigma factor activity is regulated, however, generally alternative sigma factors are retained in an inactive state by sequestration into a complex with an anti-σ factor. Upon a specific stimulus, the inhibitory effect of the anti-σ factor is alleviated and the sigma factor released for interaction with the RNA polymerase.
New insights on sigma factor activation
The team identified an ECF σ factor / threonine-kinase pair (named EcfP / PknT) that is responsible for sensing polymyxin antibiotics stress and mediating bacterial resistance towards polymyxin in the human pathogen Vibrio parahaemolyticus. Polymyxins constitute a class of commercially available antibiotics that disrupts the outer and inner membrane, resulting in cell death. Usually they are used as a last resort to treat Gram-negative infections.
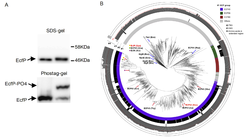
Upon treatment of V. parahaemolyticus with polymyxin antibiotics, PknT is activated and phosphorylates the σ factor EcfP. This results in EcfP activation and expression of an essential polymyxin resistance regulon. EcfP phosphorylation occurs at a highly conserved threonine residue, Thr63, which is positioned within a divergent region linking the two structural domains of EcfP, σ2.1 and σ2.2. The intrinsic inactive state of EcfP is due to the absence of a negatively charged DAED motif in the region between σ2.1 and σ2.2, which usually mediates binding to the RNAP core enzyme. Instead, the DAED motif has been replaced by non-charged amino acids, which renders EcfP inactive and unable to bind the RNAP. Strikingly, phosphorylation at residue Thr63 mimics the negative charge, usually provided by the DAED motif, and permits interaction of EcfP with RNAP in formation of the holyenzyme and the consequential expression of EcfP target genes required for polymyxin resistance.
The findings have revealed a previously unknown mechanism of transcriptional regulation, which instead relies on intrinsically inactive σ factor that is unable to bind the RNAP core enzyme. Only upon phosphorylation on a specific residue is the σ factor activated and able to bind the RNAP in formation of the holoenzyme and consequently drive expression of specific genes.
Signal transduction and transcriptional regulation act as joint forces
An extensive bioinformatics analysis indicated that transcriptional regulation by σ-phosphorylation is a general and widespread mechanism in bacteria, presenting a new paradigm in transcriptional regulation. Altogether, our discoveries reveal how nature has merged two distinct regulation mechanisms – STK signaling and regulation of σ factor activity – in order to achieve the ability of environmental adaptation. Importantly, our results indicate that this likely constitutes a widespread signaling mechanism in bacteria where the input and output cues are modular based on the sensing domain fused to the kinase and the promoter sequence recognized by the σ factors, respectively, while the signal transduction pathway of σ phosphorylation is conserved.
Unraveling the underpinnings of transcriptional regulation by σ factor phosphorylation, the cellular response to changes in the environment and how these mechanisms combined regulate stress responses, such as the antibiotic resistance in Vibrio parahaemolyticus, addresses issues that are important in general for understanding regulation of gene expression and cellular adaptation throughout the bacterial kingdom.












