It`s All About the Pole Position
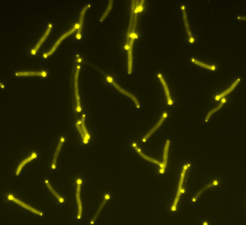
M. xanthus is a predatory soil bacterium that, when there is abundant food, forms a single species spreading colony, a so-called biofilm. Within a colony, several fascinating social interactions can be observed such as swarm formation, hunting for prey and, when the colony is starving, the formation of spore-filled fruiting bodies. This very special lifestyle makes M. xanthus a wonderful model organism to unravel the mechanisms that allow bacteria to adapt and differentiate in response to changes in the environment.
The research group of Lotte Søgaard-Andersen at the Max Planck Institute in Marburg aims to understand how M. xanthus respond to starvation within formation of spore-filled fruiting bodies. Motility is essential for M. xanthuscells to swarm and to form fruiting bodies. M. xanthus cells are rod-shaped and have two motility systems, one for type IV pili-dependent motility and one for gliding. These two motility machineries are polarized and only assemble at the leading cell pole. When M. xanthus cells reverse their direction of movement, the two motility machineries disassemble at the old leading pole and then reassemble at the new leading cell pole.
Dr. Dobromir Szadkowski in the Søgaard-Andersen lab addressed how the motility machineries assemble at the leading pole in his research. “When we started this work, we knew that this is regulated by at least three proteins that are all localized to the cell poles: MglA is a small Ras-like GTPase, which stimulates motility in its GTP-bound form. MglB is a GTPase activating protein (GAP) of MglA. We also knew that RomR is important for polar localization of MglA-GTP. MglA-GTP localizes to the leading cell pole while MglB and RomR are asymmetrically localized to the cell poles but mostly at the lagging cell pole” explains Dr. Szadkowski.
The inspiration for the new work was the observation that small GTPases in eukaryotes not only depend on a GAP for efficient GTP hydrolysis but also on a guanine nucleotide exchange factor (GEF) to efficiently exchange GDP for GTP. Dr. Szadkowski explains “So, we went searching for a GEF for MglA and this search led us to the identification of the protein that we now call RomX”. As expected for a GEF of MglA, RomX is essential for motility. It was also found that RomX localizes to the cell poles and this localization depends on RomR. The culmination of all these efforts were the finding that the RomR/RomX complex has GEF activity and brings about the polar localization of MglA-GTP.
Putting all the data together, the scientists have suggested that in an M. xanthus cell, RomR recruits RomX to the cell poles and the RomR/RomX complex, in turn, recruits MglA-GTP to the leading cell pole via two mechanisms. First RomR/RomX stimulates accumulation of MglA-GTP using its GEF activity; and, second, RomR/RomX binds MglA-GTP at the leading cell pole. Both activities contribute to a high local concentration of MglA-GTP at this pole with the stimulation of motility. MglA-GTP is kept away from the lagging cell pole because MglB with its GAP activity localizes to this pole. So, the spatially separated activities of the RomR/RomX GEF complex and the MglB GAP at opposite poles help to establish front-rear polarity for motility. The RomR/RomX complex is the first bacterial nucleotide exchange factor shown to be involved in regulating bacterial cell polarity.
Interestingly, the GEF and GAP of MglA have a spatial organization that is strikingly similar to the GEFs and GAPs of small GTPases involved in regulating cell polarity and motility in eukaryotes. The scientists also note that MglA is found widespread in prokaryotes; however, RomR and RomX have a much more limited distribution suggesting that that there might be other GEFs acting on MglA-like proteins.
This work was financially supported by the Deutsche Forschungsgemeinschaft (DFG) within the framework of the Transregio 174 “Spatiotemporal dynamics of bacterial cells” as well as by the Max Planck Society.






![<p>Discovery of [Fe]-hydrogenase in bacteria opens new possibilities for conversion of hydrogen</p>](/506336/original-1555497672.jpg?t=eyJ3aWR0aCI6MzYwLCJoZWlnaHQiOjI0MCwiZml0IjoiY3JvcCIsImZpbGVfZXh0ZW5zaW9uIjoianBnIiwib2JqX2lkIjo1MDYzMzZ9--d3d7c57e064104c2acc573916156127d6bedd85b)



![Protection of [Fe]-hydrogenase](/416905/teaser-1561027823.jpg?t=eyJ3aWR0aCI6MzYwLCJoZWlnaHQiOjI0MCwiZml0IjoiY3JvcCIsImZpbGVfZXh0ZW5zaW9uIjoianBnIiwib2JqX2lkIjo0MTY5MDV9--6ea496a03c86ba3d98a19e05aac63809917fe800)

