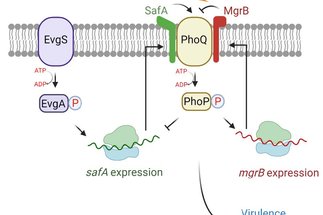
Small proteins in bacterial signaling networks
Dr. Jing Yuan
Research Area
We are interested in how bacteria sense and infect mammalian host with an emphasis on small proteins. Small proteins are involved in various biological processes, such as stress response, nutrient transport, photosynthesis, drug efflux, bacterial defense systems, cell-cell communication, and immuno-surveillance. One of the medical relevant functions of small proteins is the regulation of bacterial pathogenicity by targeting two-component signalling systems.
Two-component signalling systems are crucial for bacteria to survive and thrive in their natural habitats. Sensor kinases are the central part of two-component signal transduction systems. Some detect the mammalian host environment, leading to the expression of genes involved in membrane modification, antibiotic resistance, and the onset of bacterial pathogenicity. One such sensor kinase is PhoQ, the master regulator of the bacterial virulence program in Enterobacteriaceae. Sensor kinases are prevalent in prokaryotes and absent in mammals, making them interesting targets for new antimicrobials.
Using molecular, cellular, and synthetic approaches in combination with computational simulations and structural studies through collaborations, we aim to investigate the function of small proteins and utilize the knowledge to engineer new antimicrobials.
We currently aim to answer the following questions:
- What are the regulation mechanisms of small proteins?
- How do small proteins regulate bacterial infection?
- Can we engineer small proteins to suppress bacterial virulence?
- How to uncover the function of newly discovered small proteins?

Recent Publications
Lazaridi, S.; Yuan, J.; Lemmin, T. (2024) Atomic insights into the signaling landscape of E. coli PhoQ Histidine Kinase from Molecular Dynamics simulations. Scientific Reports, https://doi.org/10.1101/2024.04.19.590235.
Jiang, S.; Celen, G.; Glatter, T.; Niederholtmeyer, H.; Yuan, J. (2024) A cell-free system for functional studies of small membrane proteins. BioRxiv, 2023.12.22.573026
Jiang, S.; Steup, L. C.; Kippnich, C.; Lazaridi, S.; Malengo, G.; Lemmin, T.; Yuan, J. (2023) The inhibitory mechanism of a small protein reveals its role in antimicrobial peptide sensing. Proceedings of the National Academy of Sciences of the United States of America 120 (41), e2309607120
Breuer, R.; Gomes-Filho, J. V.; Yuan, J.; Randau, L. (2023) Transcriptome profiling of Nudix hydrolase gene deletions in the thermoacidophilic archaeon Sulfolobus acidocaldarius. Frontiers in Microbiology 14, 1197877
Yadavalli, S. S.; Yuan, J. (2021) Bacterial small membrane proteins: the Swiss army knife of regulators at the lipid bilayer. Journal of Bacteriology 204 (1), e0034421
Yadavalli, S. S.; Goh, T.; Carey, J. N.; Malengo, G.; Vellappan, S.; Nickels, B. E.; Sourjik, V.; Goulian, M.; Yuan, J. (2020) Functional Determinants of a Small Protein Controlling a Broadly Conserved Bacterial Sensor Kinase. Journal of Bacteriology 202 (16), e00305-20
Yuan, J.; Jin, F.; Glatter, T.; Sourjik, V. (2017) Osmosensing by the bacterial PhoQ/PhoP two-component system. Proceedings of the National Academy of Sciences of the United States of America 114 (50), pp. E10792 - E10798