Swimming bacteria shape their community
Swimming bacteria entraining non-swimmers determine the structure of mixed microbial assemblies
Bacterial communities are of great importance to humans. But how does the complex structure that is crucial for the functioning of the species form? As a team of researchers led by Dr Remy Colin from the Max Planck Institute in Marburg has discovered, purely physical interactions between swimming and non-swimming bacteria are in principle sufficient.
In laboratories, bacteria are often kept in pure culture. In the natural environment, however, they live together in complex and well-structured bacterial communities. These play a major role in the many aspects of human life that are affected by bacteria - for example, in health (gut or dental plaque microbiomes, but also medical equipment contaminations), in agriculture (plant root microbes) or in industrial processes (like bacterial floccules that process waste in water treatment plants).
The spatial structure that the different species adopt is increasingly recognized as a crucial element for them to function well together within these communities. Due to their complexity, however, the development and dynamics of these structures are poorly understood. To date, research into complex microbial communities has mainly focused on the biochemical interactions between community members. In contrast, the role of their physical interactions, particularly those resulting from swimming motility, which is common in bacteria, in structuring these communities remains unclear.
A small number of swimmers and a solid surface are all that is needed
Dr Remy Colin and his team in Prof. Dr Victor Sourjik's department at the Max Planck Institute for Terrestrial Microbiology are interested in the biophysics of motile microorganisms. Looking at the physical interactions in multi-species bacterial communities, the researchers discovered that bacteria that are able to swim induce the formation of some spatial structures. The swimmers drive the formation of large-scale patterns by other (non-swimming) bacteria through purely physical interactions. This happens close to a solid surface, where the swimmers accumulate and swim in circles.
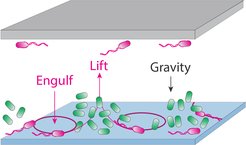
"The circular swimmers induce convective fluid flows that try to carry the non-swimmers around, a bit like pasta in boiling water, but because of their relatively strong sedimentation rate, it results in neighbouring regions of bacterial accumulation and depletion that create the patterns. Cell-cell attachment then produces structured aggregates that cement these otherwise dynamic patterns into a relatively static architecture", says Silvia Espada Burriel, PhD student and first author of the study. The researchers made their discovery through a combination of microscopy experiments and numerical simulations.
A new perspective on the organization of microbial communities
Remarkably, this mechanism requires only a fraction of the bacteria to swim and a surface. Although the researchers studied it in a well-controlled model system, it is very likely to occur in a wide variety of natural situations. "Our finding also opens up a new perspective on the organization of microbial communities," says Dr Remy Colin. "Indeed, most research to date has focused on the role of interactions mediated by specific chemicals that the bacteria produce, consume and/or exchange, whereas here we find a general, purely physical effect that may be essential for the production of structures."
While the findings are an interesting step towards a better understanding of the architecture and function of microbial communities, the researchers are far from the end of the journey, as Remy Colin points out. "Important questions that remain open and that we are currently investigating include how this physical mechanism interacts with chemical interactions in realistic microbial communities, or how this mechanism interacts with external constraints on the community, such as the presence of large-scale flows, as is the case in pipes, the gut, or the ocean."










