
Yeast and Aging
As yeast is proliferating in an unsymmetrical way, there are two different approaches to determining its age: replicative life span (RLS) and chronological life span (CLS) [8]. The chronological lifespan (CLS) measures the time a non-dividing cell can survive and keep its ability to return to a dividing stage. The replicative lifespan (RLS) reflects the number of daughter cells that are budding of one mother cell, and thus resembles the aging of mammalian cells that undergo a fixed number of divisions before dying (such as fibroblasts).
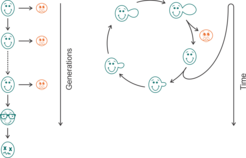
Relevance of yeast age in Research
Relevance of yeast age in biotech production
Relevance of yeast age drug develpoment
While there are technical approaches to determine the CLS in high-throughput, the RLS is in most labs still analyzed using a manual method that was already introduced in 1959.
Manual RLS determination
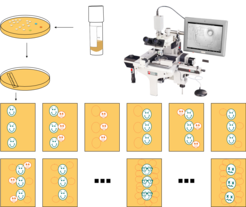
To determine the RLS manually, the yeast strain of interest is streaked from a frozen stock onto an agar plate.
After colonies have formed, they are used to inoculate a fresh plate containing the same growth medium. This main cultivation plate is incubated until colony formation can be detected.
About 40 – 60 single cells are picked from the patch of cells using a micromanipulation microscope and transferred to defined coordinates on the same plate. The plate is further incubated until the isolated cells divide for the first time after they have been transferred.
Mother and daughter cells are separated from each other using the micromanipulator again, and while the daughter cell remains at the initial coordinates of its mother, the mother cell is removed and discarded.
The number of daughter cells deriving from these “virgin” cells counted by incubating the plates for intervals of 90 – 120 minutes, interrupting the incubation, separating the cells from their own daughter cell, discarding this new daughter cell, and continuing the incubation.
In classic RLS determinations using microdissection, budding events are recorded manually and the data is digitalized for further analysis. Afterward, the data is converted into survival curves that visualize the number of budding events as relative viability
The cost of manual RLS determination
Considering that a typical RLS of a yeast strain could range from 20 – 60 divisions, on average an experiment can run over 15 days, where every 60-90 minutes a manual intervention is needed for separating the mother and daughter cells for continuous tracking of mother cells. This implies that the physical presence of a researcher is necessary for the manual isolation of daughter and mother cells. During an eight-hour workday, the procedure described above is repeated between four and six times, before the plates are stored at 4 °C to slow down the cell division overnight. This stop in the evening and start again in the morning protocol has inherent challenges i.e. the yeast could divide in the night and the entire experiment could be ruined. The comparison of two strains (e.g. healthy versus disease model, or treated versus untreated) with the minimal number of replicates to generate statistically interpretable data occupies a scientist or research assistant for a minimum of two weeks.
Research labs that focus on the use of RLS employ high numbers of student research assistants that are organized in shifts, to cover 24 h and to avoid interruptions in the cell division.
It is obvious, that this high price limits the number of experiments performed per year, apart from the fact, that the respective researcher cannot do anything else while sitting in front of a microscope, doing highly repetitive work.
Microfluidic approaches to determine the RLS in yeast
The idea to overcome the limitations of RLS determination employing microfluidics is not innovative as such. Since 2012 several research groups have published microfluidic-based cultivation devices, which allow the measurement of the RLS with a (compared to the manual method described above) increased throughput.
The devices developed for yeast aging feature clever geometrical structures to mechanically trap mother cells. While the mother cell is trapped, the offspring is flushed away and can be quantified based on a time-lapsed microscopic video after the experiment is over.
What all published methods do have in common is the requirement for sophisticated microfluidic equipment (microfluidic pumps, high-resolution microscopes, microfluidics incubators, visualization software, etc.) to use the otherwise sound microfluidic chips.
Apart from the financial point of view, designers and applicants of microfluidic chips for yeast aging studies have so far failed to prove that the results obtained using their respective devices recapitulate the results of microdissection assays. This is of particular importance, in cases where the yeast model was found to resemble the behavior of multicellular eukaryotes (calorie restriction, response to conserved genetic modifiers of aging such as sirtuins, and the target of rapamycin).
Drawbacks of current microfluidics devices for RLS determination:
Require expensive microfluidic equipment (pumps, incubators, chips)
Require expensive motorized microscopes
Require capacity to store and analyze huge amounts of video/imaging data
Are difficult to adapt to the laboratory without microfluidic expertise
Thus, there is an imminent need for an automated, simplified, stand-alone, high throughput, and integrated system for lifespan determination in yeast overcoming the highlighted limitations.

