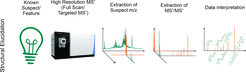
Structural Elucidation
The structural identification of a target molecule is a frequent question in analytical science. In general, it is done by aquireing as much and as accurate data about the molecule as possible (accurate mass, adduct formation, natural isotopic labelling, fragmentation, retention time using different columns,…), and compare this data to existing compound libraries or in-silico predictions of the respective parameters. The complexity of the endeavor and the methodological approach can be very different and are highly dependent on the prior knowledge a researcher has about the sample and the analytical target. If a molecule of known structure is expected to be found in a sample, the analytical task shifts from its identification to the conformation of its identity. In this case, even the chromatographic conditions can be chosen to facilitate its detection, and a high level of confidents can be achieved by matching the measurement data against online or even in-house libraries.
The less knowledge about a compound is available, the more challenging the structural elucidation gets. If the structure of a previously unknown compound should be determined, the identification can in some cases be based on an assumed structure, which is rationalized by the metabolism, chemical reactions, or the known compounds present in a sample. In this cases, confirming or contradiction arguments for the assumed structure are deduced from the prediction of the parameters determined by the MS and from the negative match against existing compound databases.
If the structure of an unknown compound that is not expected to be present in a sample should be determined, the selection of the compounds to be structurally elucidated ads another layer of complexity. In this cases, we combine an untargeted comparative approach with the established workflows for structural elucidation.
We use state-of-the-art hardware (Orbitrap ID-X) and software tools (Trace Finder, Compound Discoverer, MSDial, MetFrag, Skyliner) for our structural elucidation projects. If you want to be Involved in the data analysis process, you are welcome to do so and get some hands-on experience on the computational aspects of metabolomics.